This vignette shows how to load and check WATLAS data.
Good to know
tools4watlas is based on data.table to be
fast and efficient. A key feature of data.table is
modification in place, where data is changed without making a copy. To
prevent this (whenever it is not desired) use the function
copy() to make a true copy of the data set. Basic knowledge
about data.table
is helpful, but not necessary, when working with
tools4watlas.
Getting data
WATLAS data can either be loaded from a local SQLite database or a remote SQL database server. To do so, first select the tags and time period for which to extract data.
Use the tags_watlas_all.xlsx file (including metadata of
all tags) or for collaborators the tags_watlas_subset.xlsx
(including a subset of metadata) to select the desired tags.
Using the example data in tools4watlas, the
tags_watlas_subset.xlsx will provide a table with the
following columns:
| Column | Description |
|---|---|
| year | Year in which the bird was caught |
| species | Species common name |
| tag | Tag ID with 4 digits |
| rings | Metal ring number |
| crc | Colour ring combination |
| release_ts | Release time stamp in CET |
| catch_location | Location where the bird was caught |
Select the desired tags and time period
# file path to the metadata
fp <- system.file(
"extdata", "tags_watlas_subset.xlsx", package = "tools4watlas"
)
# load meta data
all_tags <- readxl::read_excel(fp, sheet = "tags_watlas_all") |>
data.table()
# subset desired tags using data.table
# (for example all tags from 2023)
tags <- all_tags[year == 2023]$tag
# time period for which data should be extracted form the database (in CET)
from <- "2023-09-21 00:00:00"
to <- "2023-09-25 00:00:00"Extract data from local SQLite file
First, the path and file name of the local SQLite database needs to
be provided. Then, a connection to the database can be established, and
the database can be queried for the selected tags and period. Here, we
will load the selected tagging data in a data.table
object.
# establish database connection
sqlite_db <- system.file(
"extdata", "watlas_example.SQLite", package = "tools4watlas"
)
con <- RSQLite::dbConnect(RSQLite::SQLite(), sqlite_db)
# load data from database
data <- atl_get_data(
tags,
tracking_time_start = from,
tracking_time_end = to,
timezone = "CET",
use_connection = con
)
# close connection
RSQLite::dbDisconnect(con)Alternatively, extract from remote SQL-database
To safely work with database credentials, one option is to store them as environmental variables in R. This allows, for example, to have shared scripts on GitHub without storing the credentials online. Ask Allert for the host, username and password and then add them in your environment as indicated below. After adding the credentials to the R-environment, restart R and access should be available and the scripts should run succesfully.
# open .Renviron to edit
file.edit("~/.Renviron")
# add variables
host = "host"
username = "username"
password = "password"
# access variables (example)
Sys.getenv("variable_name")Connecting to a remote database with atl_get_data() is
similar to connecting with a local SQLite database. In this example
(chunk not run and only shown), we load the last three days of data from
all tags of 2024. Note that the host, username and password are
specified as environmental variables in this example, but can also be
specified directly.
Connecting to the remote database is normally restricted to current
group members that also have access to the NIOZ/Bijleveld Teams
environment. To use the most recent metadata stored online, we first
load the tags_watlas_all.xlsx from the “WATLAS” SharePoint
folder: Documents/data/. Either specify the path to your
local copy of this folder or add the path for your user in the
atl_file_path() function.
# file path to WATLAS teams data folder
fp <- atl_file_path("watlas_teams")
# load meta data
all_tags <- readxl::read_excel(
paste0(fp, "tags/tags_watlas_all.xlsx"),
sheet = "tags_watlas_all"
) |>
data.table()
# subset all tags from 2024
tags <- all_tags[year == 2024]$tag
# select N last days to get data from
days <- 3
from <- (Sys.time() - 86400 * days) |> as.character()
to <- (Sys.time() + 3600) |> as.character()
# load data from database
data <- atl_get_data(
tags,
tracking_time_start = from,
tracking_time_end = to,
timezone = "CET",
host = Sys.getenv("host"),
database = "atlas2024",
username = Sys.getenv("username"),
password = Sys.getenv("password")
)Data explanation
The resulting loaded WATLAS data will be a data.table
with the following columns:
| posID | tag | time | datetime | x | y | nbs | varx | vary | covxy |
|---|---|---|---|---|---|---|---|---|---|
| 1 | 3027 | 1695438802 | 2023-09-23 03:13:22 | 650692.8 | 5902549 | 3 | 46.97 | 392.42 | 126.62 |
| 2 | 3027 | 1695438805 | 2023-09-23 03:13:25 | 650705.6 | 5902576 | 3 | 49.09 | 460.84 | 141.21 |
| 3 | 3027 | 1695438808 | 2023-09-23 03:13:28 | 650691.6 | 5902536 | 3 | 58.18 | 471.81 | 155.89 |
| 4 | 3027 | 1695439189 | 2023-09-23 03:19:49 | 650728.6 | 5902571 | 3 | 49.97 | 441.46 | 138.20 |
| 5 | 3027 | 1695439192 | 2023-09-23 03:19:52 | 650721.0 | 5902556 | 3 | 5.34 | 28.16 | 8.58 |
| 6 | 3027 | 1695439195 | 2023-09-23 03:19:55 | 650721.1 | 5902559 | 3 | 5.55 | 35.03 | 10.43 |
| Column | Description |
|---|---|
| posID | Unique identifier for positions |
| tag | 4 digit tag number (character), i.e. last 4 digits of the full tag number |
| time | UNIX time (seconds) |
| datetime | Datetime in POSIXct (UTC) |
| x | X-coordinates in meters (UTM 31 N) |
| y | Y-coordinates in meters (UTM 31 N) |
| nbs | Number of Base Stations used for estimating the positions |
| varx | Variance in estimating X-coordinates |
| vary | Variance in estimating Y-coordinates |
| covxy | Co-variance between X- and Y-coordinates |
Remove data before release
Because tags are turned on before the bird are released, these
positions need to be removed. The release time stamp is specified in the
metadata that was previously loaded in the object
all_tags.
# correct time zone to CET and change to UTC
all_tags[, release_ts := force_tz(as_datetime(release_ts), tzone = "CET")]
all_tags[, release_ts := with_tz(release_ts, tzone = "UTC")]
# join release_ts with data
all_tags[, tag := as.character(tag)]
data[all_tags, on = "tag", `:=`(release_ts = i.release_ts)]
# exclude positions before the release
data <- data[datetime > release_ts]
# remove release_ts column from data
data[, release_ts := NULL]Add species column (or other relevant columns)
If we are working with multiple species, then we can join the species
from the metadata. In this case, species is added as the
first column of the data.table.
# join with species data
all_tags[, tag := as.character(tag)]
data[all_tags, on = "tag", `:=`(species = i.species)]
# make species first column
setcolorder(data, c("species", setdiff(names(data), c("species"))))
# order data.table
setorder(data, species, tag, time)We can also add other metadata by merging any column (e.g. color rings and catch location). However, when working with large data sets, it is advised to only add columns that are necessary. Adding columns can be done at any stage of the analyses. Here, after showing how to add columns, we delete them immediately because they are not necessary.
# join with metal rings, color rings and catch location
all_tags[, tag := as.character(tag)]
data[all_tags, on = "tag", `:=`(
rings = i.rings,
crc = i.crc,
catch_location = i.catch_location
)]
# delete columns
data[, c("rings", "crc", "catch_location") := NULL]Save data
At this point it might be good to save the raw data for further
analyses, as extracting the data from the database can take a long time
with big datasets. A convenient and fast way is to use
fwrite() from the data.table package. By
including yaml = TRUE we make sure the data stays in the
same format, when we load it again with
fread(filepath, yaml = TRUE). Note that the file path needs
to be changed when running this example.
# save data
fwrite(data, file = "../inst/extdata/watlas_data_raw.csv", yaml = TRUE)Check data
Data summary
Here, we inspect for how many individuals we have data within the selection, and how many positions we have per tag and date.
# load data
data <- fread("../inst/extdata/watlas_data_raw.csv", yaml = TRUE)
# data summary
data_summary <- atl_summary(data, id_columns = c("species", "tag"))
# N individuals with tagging data
data_summary |> nrow()## [1] 8
# N by species
data_summary[, .N, by = species]## species N
## <char> <int>
## 1: bar-tailed godwit 1
## 2: curlew 1
## 3: dunlin 1
## 4: oystercatcher 1
## 5: red knot 1
## 6: redshank 1
## 7: sanderling 1
## 8: turnstone 1
# show head of the table
data_summary |> knitr::kable(digits = 2)| species | tag | n_positions | first_data | last_data | days_data | min_gap | max_gap | max_gap_factor | fix_rate |
|---|---|---|---|---|---|---|---|---|---|
| bar-tailed godwit | 3063 | 12614 | 2023-09-23 03:27:49 | 2023-09-23 22:24:55 | 0.8 | 3 | 2541 | 42.4 min | 0.55 |
| curlew | 3100 | 8423 | 2023-09-23 04:21:46 | 2023-09-23 21:41:16 | 0.7 | 3 | 16145 | 4.5 hours | 0.41 |
| dunlin | 3212 | 3856 | 2023-09-23 00:00:00 | 2023-09-23 23:59:56 | 1.0 | 8 | 4784 | 1.3 hours | 0.36 |
| oystercatcher | 3158 | 12533 | 2023-09-23 00:00:01 | 2023-09-23 23:59:57 | 1.0 | 3 | 3060 | 51 min | 0.44 |
| red knot | 3038 | 15959 | 2023-09-23 00:00:01 | 2023-09-23 23:59:57 | 1.0 | 3 | 2850 | 47.5 min | 0.55 |
| redshank | 3027 | 15859 | 2023-09-23 03:13:22 | 2023-09-23 22:24:26 | 0.8 | 3 | 2523 | 42 min | 0.69 |
| sanderling | 3288 | 8131 | 2023-09-23 00:00:03 | 2023-09-23 23:59:54 | 1.0 | 6 | 2622 | 43.7 min | 0.56 |
| turnstone | 3188 | 10195 | 2023-09-23 00:00:45 | 2023-09-23 23:41:50 | 1.0 | 3 | 5742 | 1.6 hours | 0.36 |
| Column | Description |
|---|---|
| species | Species |
| tag | Tag number |
| n_positions | Number of positions |
| first_data | Datetime of first position (UTC) |
| last_data | Datetime of last position (UTC) |
| days_data | Days of data |
| min_gap | Minimum time interval between positions (should be the interval of the tag in seconds) |
| max_gap | Maximum time interval between positions (largest gap in seconds) |
| max_gap_factor | Maximum time interval between positions as factor (in seconds, minutes, hours, or days) |
| fix_rate | The average fix rate (=1 if every min_gap has a
position between first_data and
last_data) |
Plot the number of positions by day. Here, we use the example data that only has one day, but this graph is particularly convenient to obtain a quick overview of the data for an entire field season.
# add date
data[, date := as.Date(datetime)] |> invisible()
# N positions by species and day
data_subset <- data[, .N, by = .(tag, date)]
# plot data
ggplot(data_subset, aes(x = date, y = tag, fill = N)) +
geom_tile() +
scale_fill_viridis(
option = "A", discrete = FALSE, trans = "log10", name = "N positions",
breaks = trans_breaks("log10", function(x) 10^x),
labels = trans_format("log10", math_format(10^.x)),
direction = -1
) +
labs(x = "Date", y = "Tag") +
theme_classic()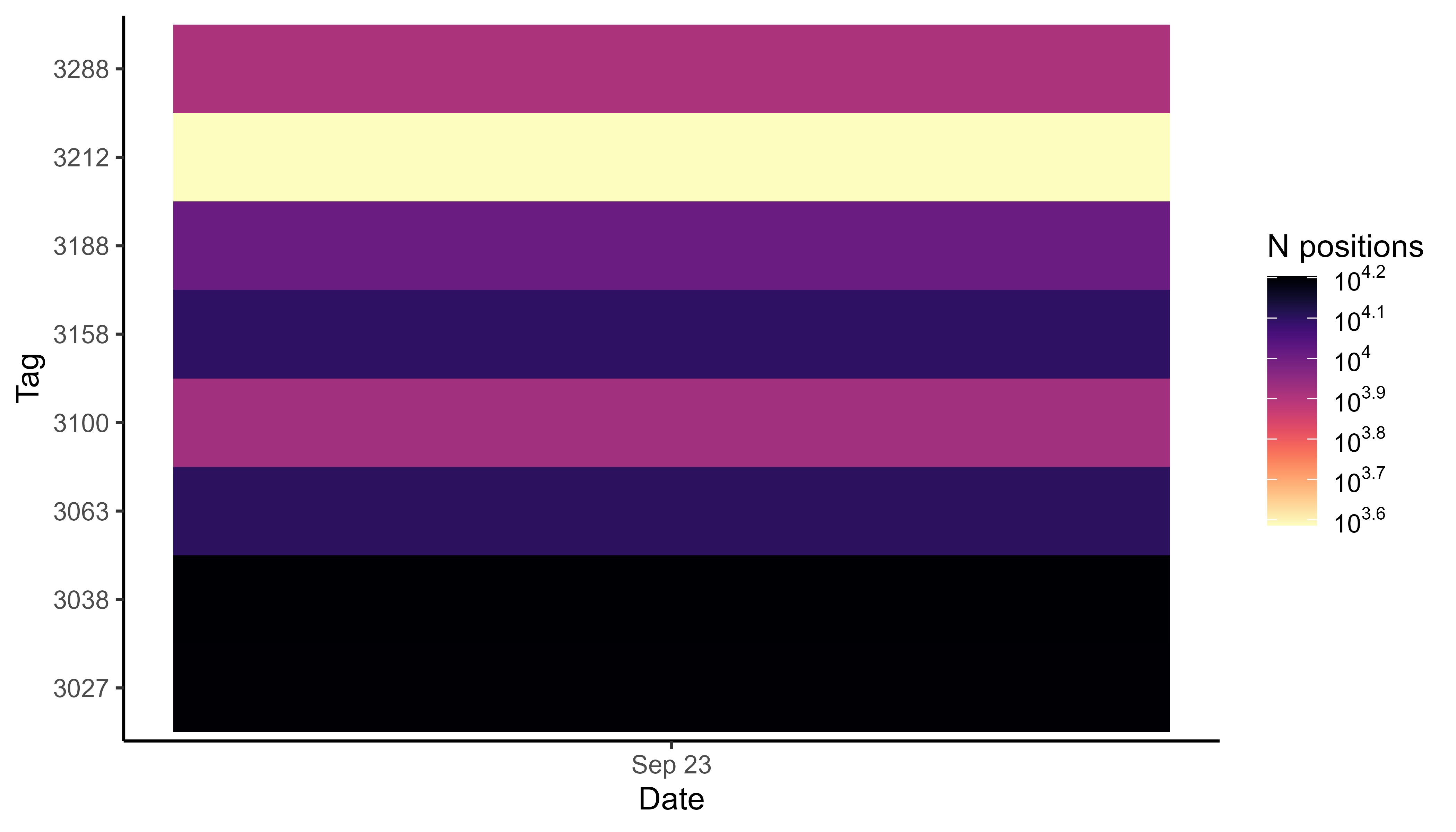
Number of positions per day by tag
Plot a heatmap of the data
With large datasets it is convenient to plot heatmaps, as plotting millions of positions is slow. For plotting all positions by tag number, for example, please see the vignette Plot data.
# create basemap
bm <- atl_create_bm(data, buffer = 800)
# round example data to 1 ha (100x100 meter) grid cells
data[, c("x_round", "y_round") := list(
plyr::round_any(x, 100),
plyr::round_any(y, 100)
)]
# N by location
data_subset <- data[, .N, by = c("x_round", "y_round")]
# plot heatmap
bm +
geom_tile(
data = data_subset, aes(x_round, y_round, fill = N),
linewidth = 0.1, show.legend = TRUE
) +
scale_fill_viridis(
option = "A", discrete = FALSE, trans = "log10", name = "N positions",
breaks = trans_breaks("log10", function(x) 10^x),
labels = trans_format("log10", math_format(10^.x)),
direction = -1
)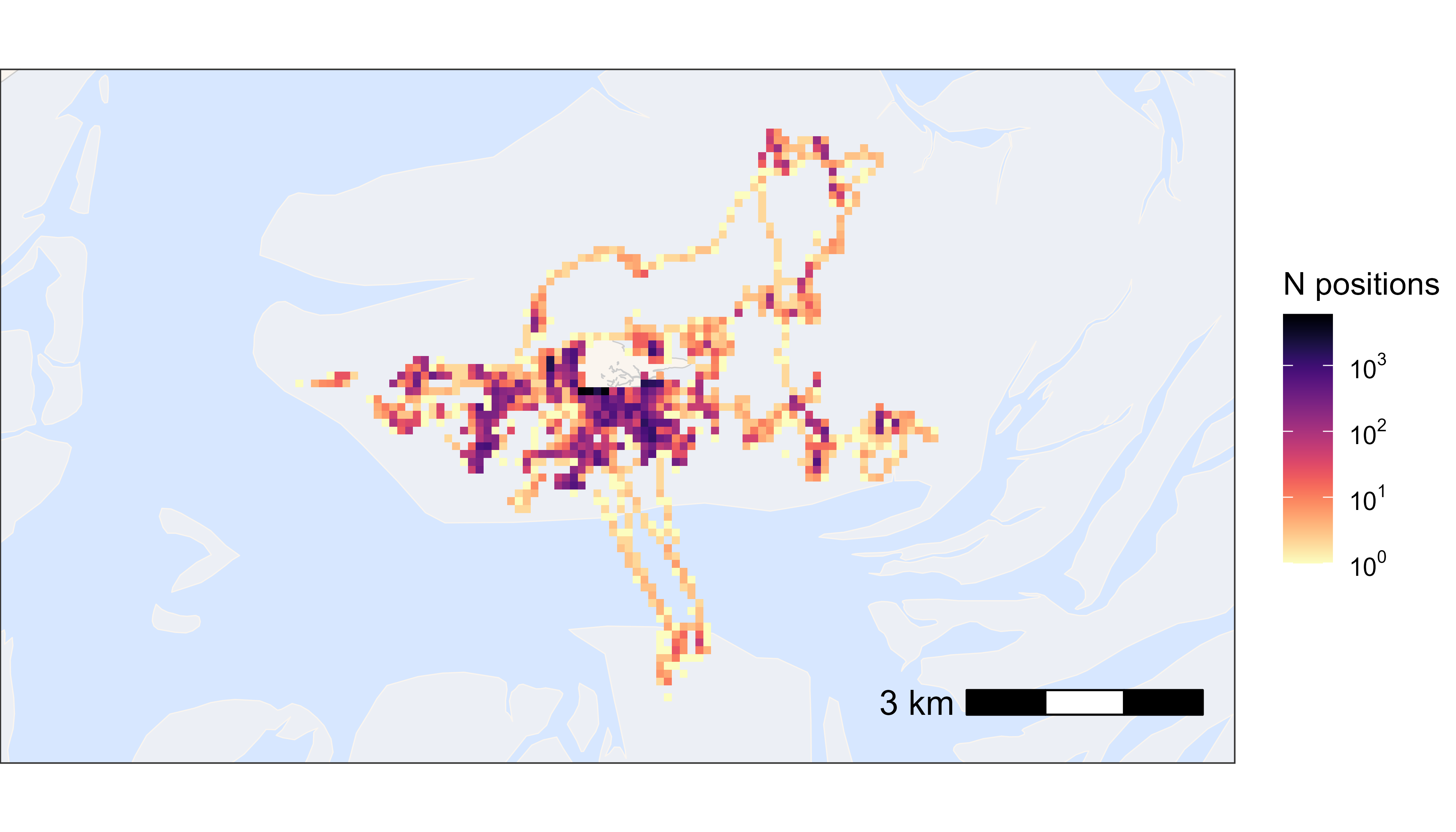
Heatmap of all positions
