Add residence patches
Source:vignettes/extended_workflow/add_residence_patches.Rmd
add_residence_patches.RmdIn this vignette, we provide a general workflow to group WATLAS position data into so-called ‘residence patches’. Note that the parameter settings should be adapted for different species’ behaviour and data quality.
Background and parameter explanation
The atl_res_patch() function is designed to segment and
aggregate WATLAS movement data into residence patches. The main
parameter is speed (max_speed). With perfect data that
would be the only parameter necessary to adjust, because the speed
flying, walking or standing do not overlap. Because WATLAS data have
localization error (comparable to GPS, see Beardsworth
et al. 2022) and gaps when birds are out of range of receivers, we
need to have additional variables for classifying these data into robust
residence patches.
The logic of the function is to first identify proto-patches
(preliminary residence patches). When subsequent positions have a speed
smaller than max_speed, a distance smaller than
lim_spat_indep and a time gap smaller than
lim_time_indep, they are assigned to the same proto patch.
Proto-patches with fewer than min_fixes positions are
filtered out.
For each proto-patch, the median position is calculated as well as
the time between two subsequent proto-patches (time between last
location and first location of the next proto-patch). Proto patches are
merged into residence patches, if the distance between the median
positions of two subsequent proto-patches is smaller than
lim_spat_indep and the time between the proto-patch is less
then lim_time_indep.
Lastly, a unique patch ID is assigned to each residence patch ordered by time from 1 to n.
Note that without cleaning the data or having short intervals (e.g. 3
sec), position error can lead to speed outliers, which affects the
creation of proto-patches. When calculating residence patches, it is
therefore recommended to first filter (var_max < 5000)
and smooth (moving_window = 5) the data. Depending on the
species’ behaviour or quality of the data, the max_speed
can also be set to a higher value, or the data can be thinned first.
Keep in mind that smoothing and thinning will influence the speeds
between positions, and thus the creation of residence patches.
Parameter overview:
-
max_speed: A numeric value specifying the maximum speed (m/s) between two subsequent positions that would be considered non-transitory. -
lim_spat_indep: A numeric value specifying the maximum distance (m) between subsequent residence patches for them to be considered independent. In combination withlim_time_indep, this parameter avoids making a new proto-patch from gaps in the data when the bird was actually still at the same location. -
lim_time_indep: A numeric value specifying the time difference (min) between two subsequent residence patches for them to be considered independent. In combination withlim_spat_indep, this parameter can prevent the creation new proto-patches when there are large gaps in the data. For example, at the roost site a bird might not move for a long time at a location with poor coverage by receivers. If the bird then moves away and sends data from the same position, we can assume that all missed positions were also at this place. -
min_fixes: The minimum number of positions for proto-patches. To make sure that residence patches have at least a few positions. -
min_duration: The minimum duration (s) for classifying residence patches. With a high-sampling interval (e.g. 1 s), short residence patches can be created, which are not biological relevant.
Guidelines to choose parameters:
In general, when deciding on the optimal parameters, the key is to find a good balance between true and false positives. We give some starting points for each parameter, which has to be adjusted based on the data quality and species behaviour.
-
max_speed: Should be as high as possible in between walking and flying speeds. A good starting point is 3 m/s, but could be reduced if it prevents the creation of proto-patches, or increased if too many proto-patches are created. Having too many proto-patches is not always an issue becasue subsequent proto-patches can be merged if the distance between them is small enough (set bylim_spat_indep). Likewise, errors in positiong data can inflate the creation of proto patches, but these will be corrected in the procedure. -
lim_spat_indep: Provides the distance between two proto-patches at which they are considered independent. This is a key parameter, because it prevents the creation of new proto-patches when the bird is still at the same location. A good starting point is 75 m, but could be increased if the position data has many/large gaps, or if the species has elongated foraging patches. -
lim_time_indep: Typically, 180 min (3 hours) works fine, but could be increased with position data that has large gaps. For example, when the analysis is focused on roosting behaviour, this could even be increased to e.g. 6 hours (360 min), to deal with large gaps in the data that can occur with roosting birds not moving and being at the same place with bad signal strength. -
min_fixes: This should be set as small as possible and 3 positions usually works. Setting this variable >1 allows assigning proto patches only if subsequent positions are consistently above the 1max_speedandlim_spat_indepand not just once becasue of an outlier, for instance. -
min_duration: Should be set as small as possible while maintaining most biological relevance. A value of 120 sec (2 minutes) seems reasonable.
Load packages and required data
# packages
library(tools4watlas)
library(ggplot2)
library(viridis)
library(foreach)
library(doFuture)
# load example data
data <- data_example
# file path to WATLAS teams data folder
fp <- atl_file_path("watlas_teams")
# load tide pattern data
tidal_pattern <- fread(paste0(
fp, "waterdata/allYears-tidalPattern-west_terschelling-UTC.csv"
))Calculate residence patches by tag
To reduce the memory size for parallel computing, we will first
subset the relevant columns from the data. This can be skipped for small
data tables. We will then run atl_res_patch() for each tag
ID in parallel. The column patch is added to the data
table, which provides the assigned patch ID’s for the positions.
# subset relevant columns
data <- data[, .(species, posID, tag, time, datetime, x, y, tideID)]
# extract the unique tag IDs
id <- unique(data$tag)
# register cores and backend for parallel processing
registerDoFuture()
plan(multisession)
# loop through all tags to calculate residence patches
data <- foreach(i = id, .combine = "rbind") %dofuture% {
atl_res_patch(
data[tag == i],
max_speed = 3, lim_spat_indep = 75, lim_time_indep = 180,
min_fixes = 3, min_duration = 120
)
}
# close parallel processing
plan(sequential)
# show head of the summary table
head(data) |> knitr::kable(digits = 2)| species | posID | tag | time | datetime | x | y | tideID | patch |
|---|---|---|---|---|---|---|---|---|
| redshank | 2 | 3027 | 1695438805 | 2023-09-23 03:13:25 | 650705.6 | 5902556 | 2023513 | 1 |
| redshank | 3 | 3027 | 1695438808 | 2023-09-23 03:13:28 | 650705.6 | 5902556 | 2023513 | 1 |
| redshank | 4 | 3027 | 1695439189 | 2023-09-23 03:19:49 | 650721.0 | 5902559 | 2023513 | 1 |
| redshank | 5 | 3027 | 1695439192 | 2023-09-23 03:19:52 | 650721.1 | 5902559 | 2023513 | 1 |
| redshank | 6 | 3027 | 1695439195 | 2023-09-23 03:19:55 | 650723.1 | 5902564 | 2023513 | 1 |
| redshank | 7 | 3027 | 1695439198 | 2023-09-23 03:19:58 | 650723.1 | 5902564 | 2023513 | 1 |
Evaluate residence patch classification and parameters
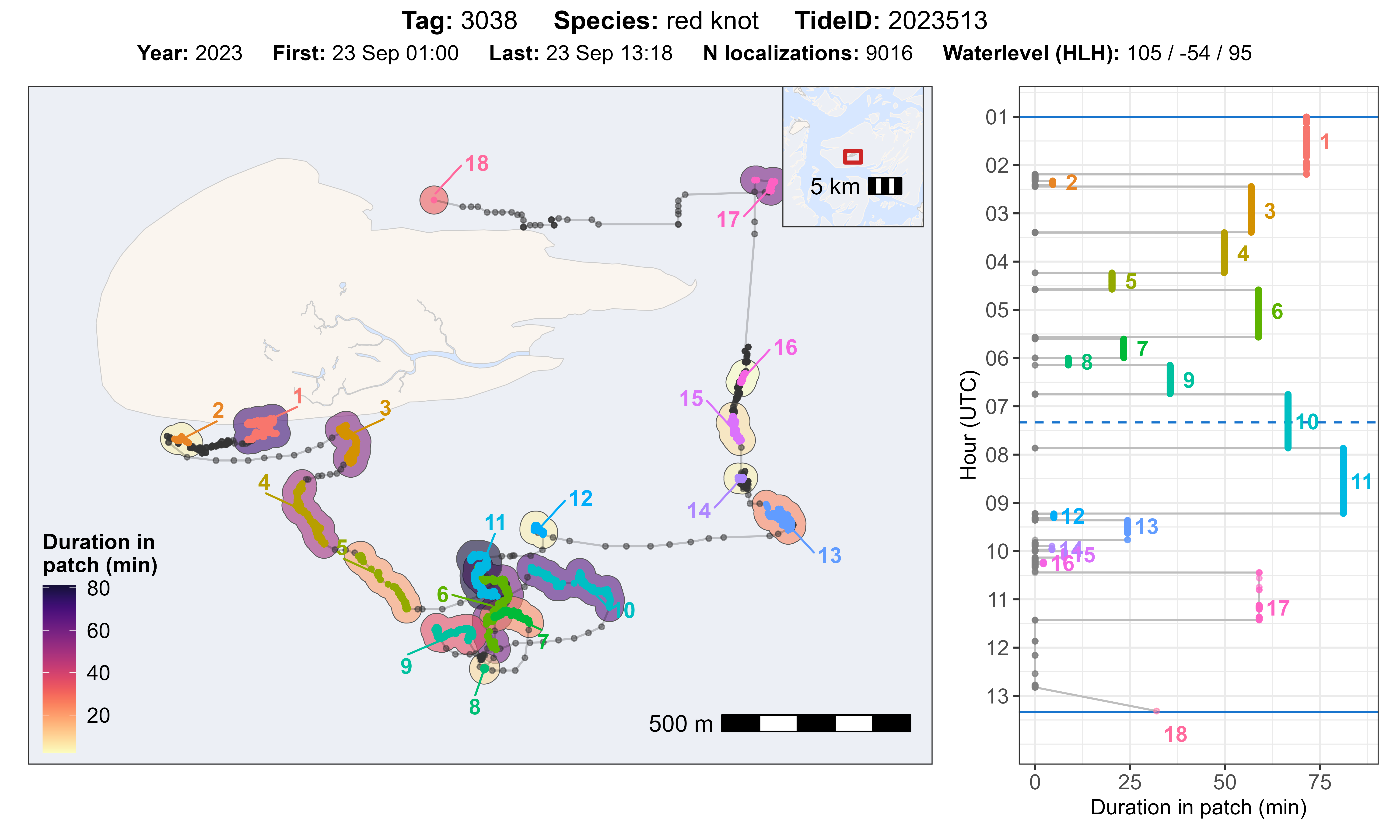
The function atl_check_res_patch() can be used to
evaluate the residence patch classification by tag and tide ID. the
function plots the track with residence patches on a map and shows the
duration (time in a patch in min) as coloured polygon on the map and as
plot against the time in a separate plot. Time starts on the top and
goes from high tide to high tide (solid blue lines), as well as
indicating low tide (dashed blue line). The title of the plot gives
basic information about the data and the water level for the
corresponding tide.
Inspect one tag and tide
We can select one tag and tide to plot. Additionally, we need to
specify the offset for the tidal data we use (e.g. 30 min for
West-Terschelling) and a buffer (in m) around the residence patch data
to create the polygon. This buffer should be set to half of
lim_spat_indep (maximum distance between subsequent
residence patches at which they will be considered independent),
ensuring that the polygons around residence patches correspond to the
spatial distance threshold used to merge residence patches.
atl_check_res_patch(
data[tag == "3038"], tide_data = tidal_pattern,
tide = "2023513", offset = 30,
buffer_res_patches = 75 / 2
)
Inspect many tags and tides
To get a general overview, we can also loop through and plot all data
by tag and tide, or for example a random sample of 100 tags and tides.
The plots are saved in any directory
(e.g. ./outputs/res_patch_check/), which has to be created
before running the code.
# create table with data combinations to plot
idc <- unique(data[, c("species", "tag", "tideID")])
# sample 100 combinations to plot
set.seed(123)
idc <- idc[sample(.N, 100)]
# register cores and backend for parallel processing
registerDoFuture()
plan(multisession)
# loop to make plots for all
foreach(i = seq_len(nrow(idc))) %dofuture% {
# plot and save for each combination
atl_check_res_patch(
data[tag == idc$tag[i]],
tide_data = tidal_pattern,
tide = idc$tideID[i], offset = 30,
buffer_res_patches = 75 / 2,
filename = paste0(
"./outputs/res_patch_check/",
idc$species[i], "_tag_", idc$tag[i], "_tide_", idc$tideID[i]
)
)
}
# close parallel processing
plan(sequential)Based on these plots and perhaps additional checks, the parameters can be adjusted to improve the classification of residence patches.
Summary of residence patch data
Once satisfied with the residence patch classification, we can summarize the residence patches by tag and patch ID and merge the desired columns back to our full data table.
# summary of residence patches
data_summary <- atl_res_patch_summary(data)
# standardise duration to minutes
data_summary[, duration := duration / 60]
# merge desired summary columns with original data table
data[data_summary, on = c("tag", "patch"), `:=`(
duration = i.duration,
disp_in_patch = i.disp_in_patch
)]
# show head of the summary table
head(data_summary) |> knitr::kable(digits = 2)| tag | patch | nfixes | x_mean | x_median | x_start | x_end | y_mean | y_median | y_start | y_end | time_mean | time_median | time_start | time_end | dist_start_end | dist_in_patch | dist_bw_patch | time_bw_patch | disp_in_patch | duration |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 3027 | 1 | 52 | 650705.5 | 650702.8 | 650705.6 | 650701.9 | 5902566 | 5902562 | 5902556 | 5902570 | 2023-09-23 03:30:41 | 2023-09-23 03:25:08 | 2023-09-23 03:13:25 | 2023-09-23 04:19:06 | 14.61 | 178.36 | NA | NA | 14.61 | 65.70 |
| 3027 | 2 | 60 | 650776.6 | 650776.6 | 650778.7 | 650771.9 | 5902216 | 5902217 | 5902216 | 5902206 | 2023-09-23 04:25:34 | 2023-09-23 04:25:32 | 2023-09-23 04:24:00 | 2023-09-23 04:27:09 | 12.29 | 51.41 | 362.21 | 293.98 | 12.29 | 3.15 |
| 3027 | 3 | 2456 | 650760.9 | 650762.0 | 650778.4 | 650699.8 | 5901722 | 5901737 | 5902014 | 5901490 | 2023-09-23 05:38:18 | 2023-09-23 05:35:44 | 2023-09-23 04:27:36 | 2023-09-23 06:51:24 | 530.22 | 1968.58 | 192.16 | 27.00 | 530.22 | 143.79 |
| 3027 | 4 | 64 | 648364.2 | 648362.5 | 648360.7 | 648365.8 | 5901596 | 5901590 | 5901578 | 5901620 | 2023-09-23 06:58:00 | 2023-09-23 06:58:01 | 2023-09-23 06:56:18 | 2023-09-23 06:59:42 | 42.01 | 79.03 | 2340.88 | 293.98 | 42.01 | 3.40 |
| 3027 | 5 | 41 | 648059.6 | 648058.4 | 648059.7 | 648058.4 | 5902193 | 5902192 | 5902184 | 5902204 | 2023-09-23 07:01:48 | 2023-09-23 07:01:51 | 2023-09-23 07:00:39 | 2023-09-23 07:02:57 | 19.69 | 48.94 | 641.75 | 57.00 | 19.69 | 2.30 |
| 3027 | 6 | 316 | 647775.8 | 647776.4 | 647771.5 | 647775.4 | 5902555 | 5902555 | 5902560 | 5902546 | 2023-09-23 07:12:32 | 2023-09-23 07:12:34 | 2023-09-23 07:03:45 | 2023-09-23 07:21:03 | 13.69 | 452.32 | 456.97 | 48.00 | 13.69 | 17.30 |
| Column | Description |
|---|---|
| tag | 4 digit tag ID (character), i.e. last 4 digits of the full tag number |
| patch | Patch ID |
| nfixes | Number of fixes in the patch |
| x_mean | Mean X-coordinate in meters (UTM 31 N) |
| x_median | Median X-coordinate in meters (UTM 31 N) |
| x_start | X-coordinate at the start of the residence patch (UTM 31 N) |
| x_end | X-coordinate at the end of the residence patch (UTM 31 N) |
| y_mean | Mean Y-coordinate in meters (UTM 31 N) |
| y_median | Median Y-coordinate in meters (UTM 31 N) |
| y_start | Y-coordinate at the start of the residence patch (UTM 31 N) |
| y_end | Y-coordinate at the end of the residence patch (UTM 31 N) |
| time_mean | Mean datetime of the positions in the residence patch |
| time_median | Median datetime of the positions in the residence patch |
| time_start | Start datetime of the patch |
| time_end | End datetime of the patch |
| dist_start_end | Distance (in meters) between first and last position |
| dist_in_patch | Distance (in meters) travelled within the patch (cumulative distance) |
| dist_bw_patch | Distance (in meters) between end of previous and start of current patch |
| time_bw_patch | Time (in seconds) between end of previous and start of current patch |
| disp_in_patch | Straight-line displacement (in meters) between start and end of the patch |
| duration | Time duration (in seconds) between first and last position in patch |
Plot the residence patches
It might also be useful to plot the residence patches using
ggplot2.
Plot by tag
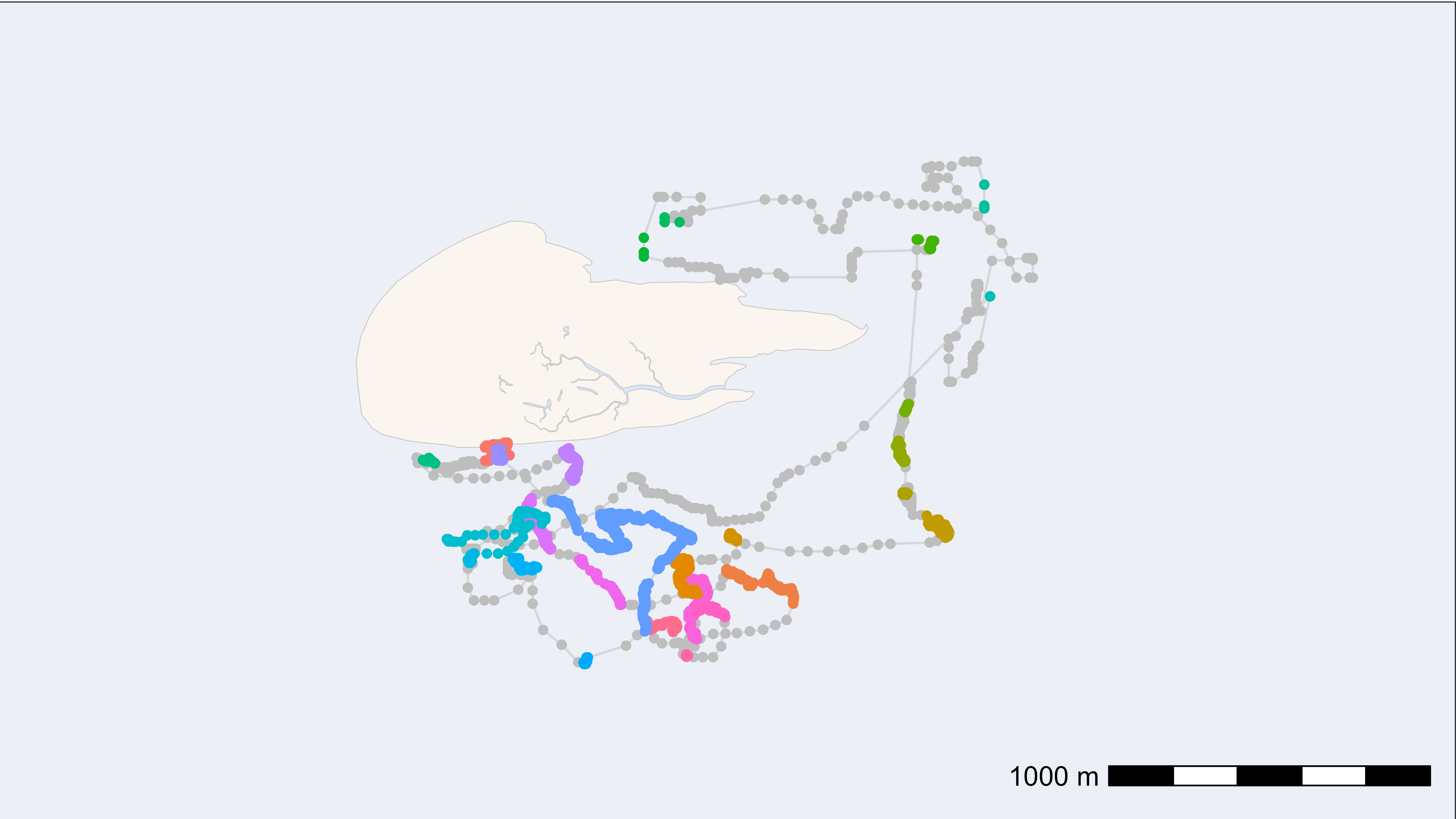
In this example, we will plot the residence patches for one red knot (tag 3038). In the frist example, the residence patches are coloured by patch ID. To show the full track, the transient (unassigned) positions are plotted in grey.
# subset red knot
data_subset <- data[tag == 3038]
data_summary_subset <- data_summary[tag == 3038]
# create basemap
bm <- atl_create_bm(data_subset, buffer = 500)
# track with residence patches coloured
bm +
geom_path(data = data_subset, aes(x, y), alpha = 0.1) +
geom_point(
data = data_subset, aes(x, y), color = "grey",
show.legend = FALSE
) +
geom_point(
data = data_subset[!is.na(patch)], aes(x, y, color = as.character(patch)),
size = 1.5, show.legend = FALSE
)
In the second example, the residence patches are plotted at their median positions with the size and colour scaled to their duration (in minutes).
# plot residence patches itself by duration
bm +
geom_point(
data = data_summary_subset,
aes(x_median, y_median, color = duration, size = duration),
show.legend = TRUE, alpha = 0.5
) +
scale_color_viridis()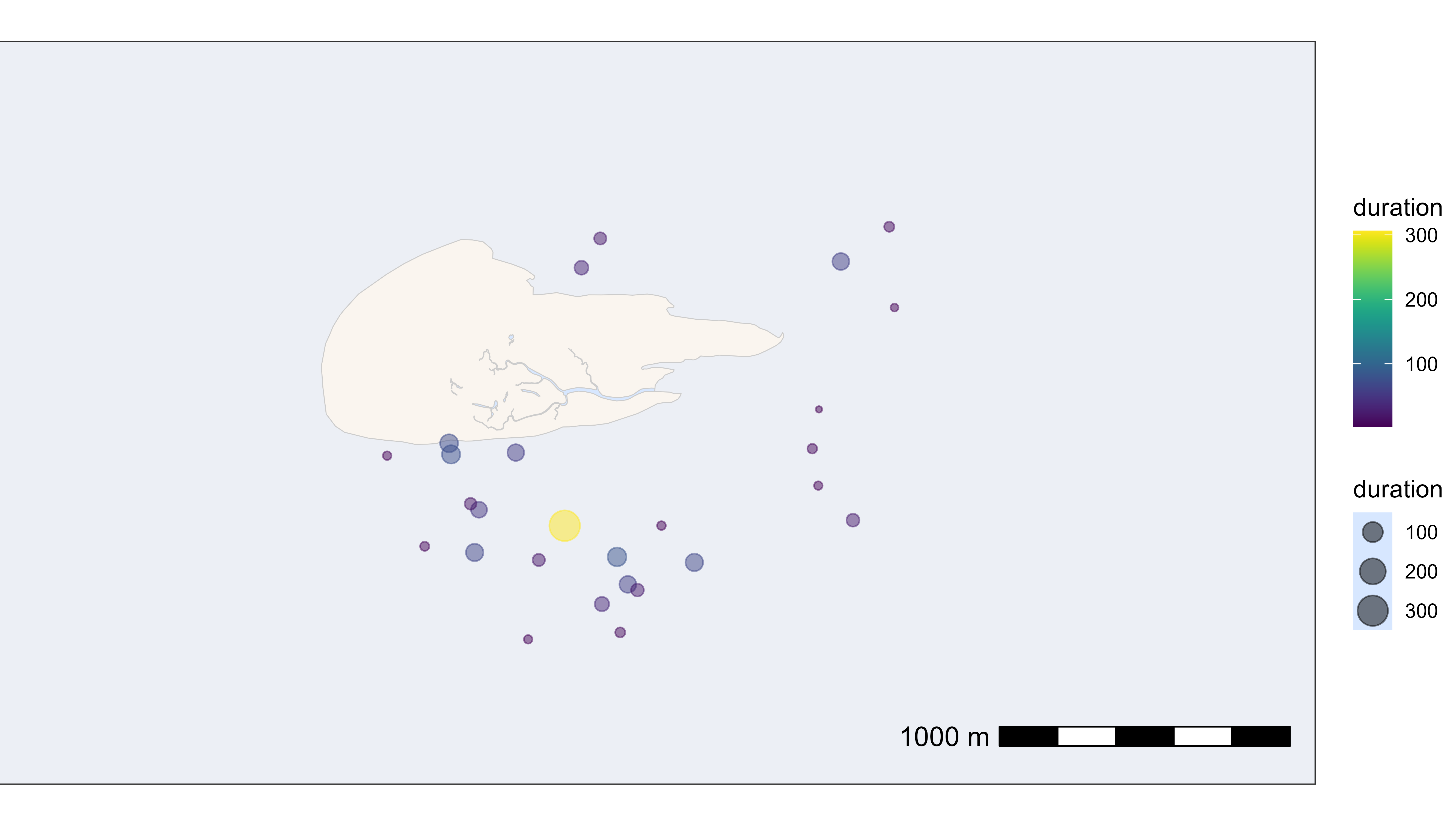
In the third example, we will calculate polygons around the residence patches and plot them
# make patch character for plotting
data_subset[, patch := as.character(patch)]
# create polygons around residence patches
d_sf <- atl_as_sf(
data_subset,
additional_cols = "patch",
option = "res_patches", buffer = 75 / 2
)
# geom_sf overwrites the coordinate system, so we need to set the limits again
bbox <- atl_bbox(data_subset, buffer = 500)
# plot polygons around residence patches
bm +
# add patch polygons
geom_sf(data = d_sf, aes(fill = patch), alpha = 0.2) +
# add track and points
geom_path(
data = data_subset, aes(x, y),
linewidth = 0.5, alpha = 0.5
) +
geom_point(
data = data_subset[is.na(patch)], aes(x, y),
size = 0.5, alpha = 0.5, color = "grey20",
show.legend = FALSE
) +
geom_point(
data = data_subset[!is.na(patch)], aes(x, y, color = patch),
size = 0.5, show.legend = FALSE
) +
# set extend again (overwritten by geom_sf)
coord_sf(
xlim = c(bbox["xmin"], bbox["xmax"]),
ylim = c(bbox["ymin"], bbox["ymax"]), expand = FALSE
)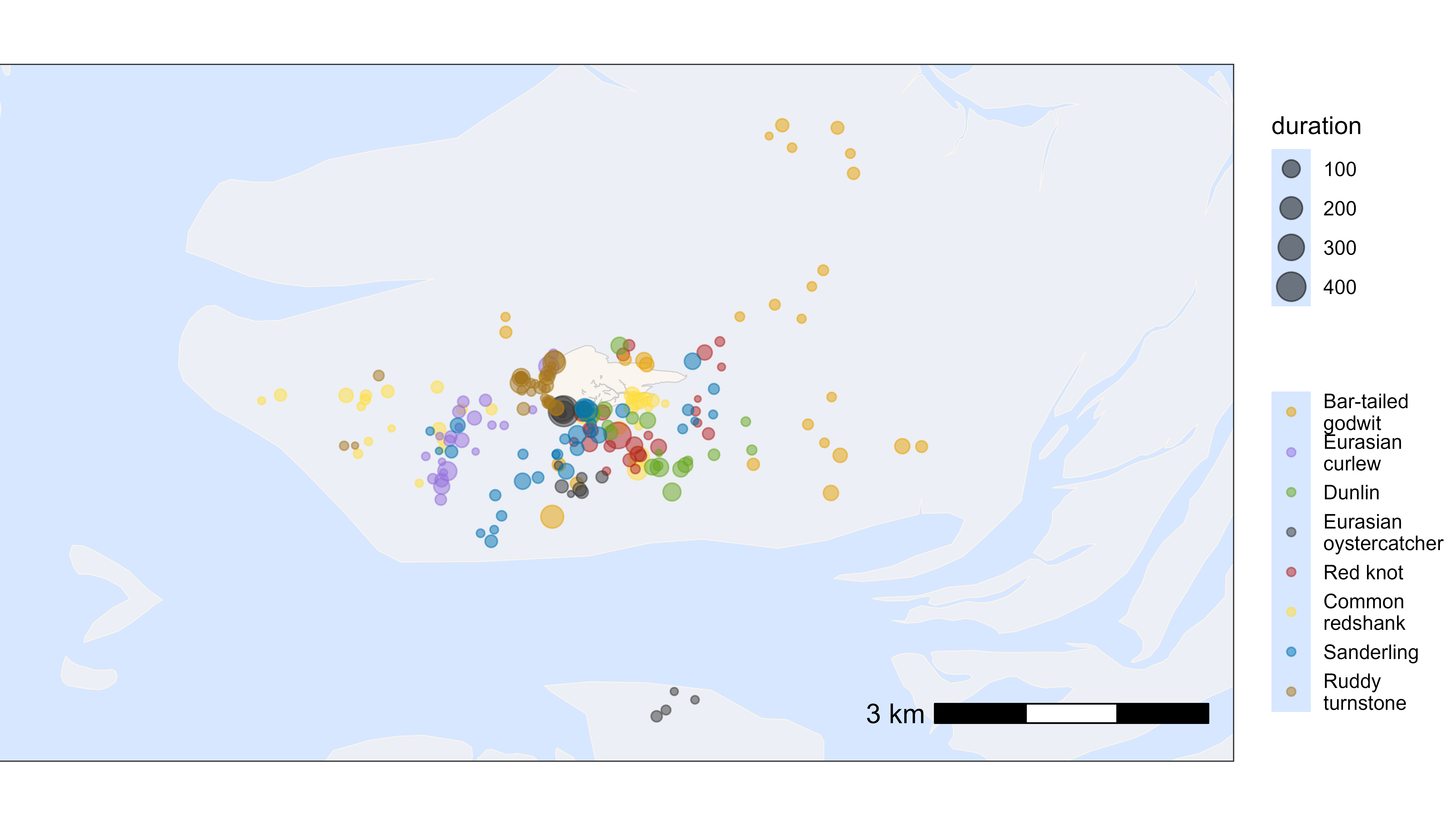
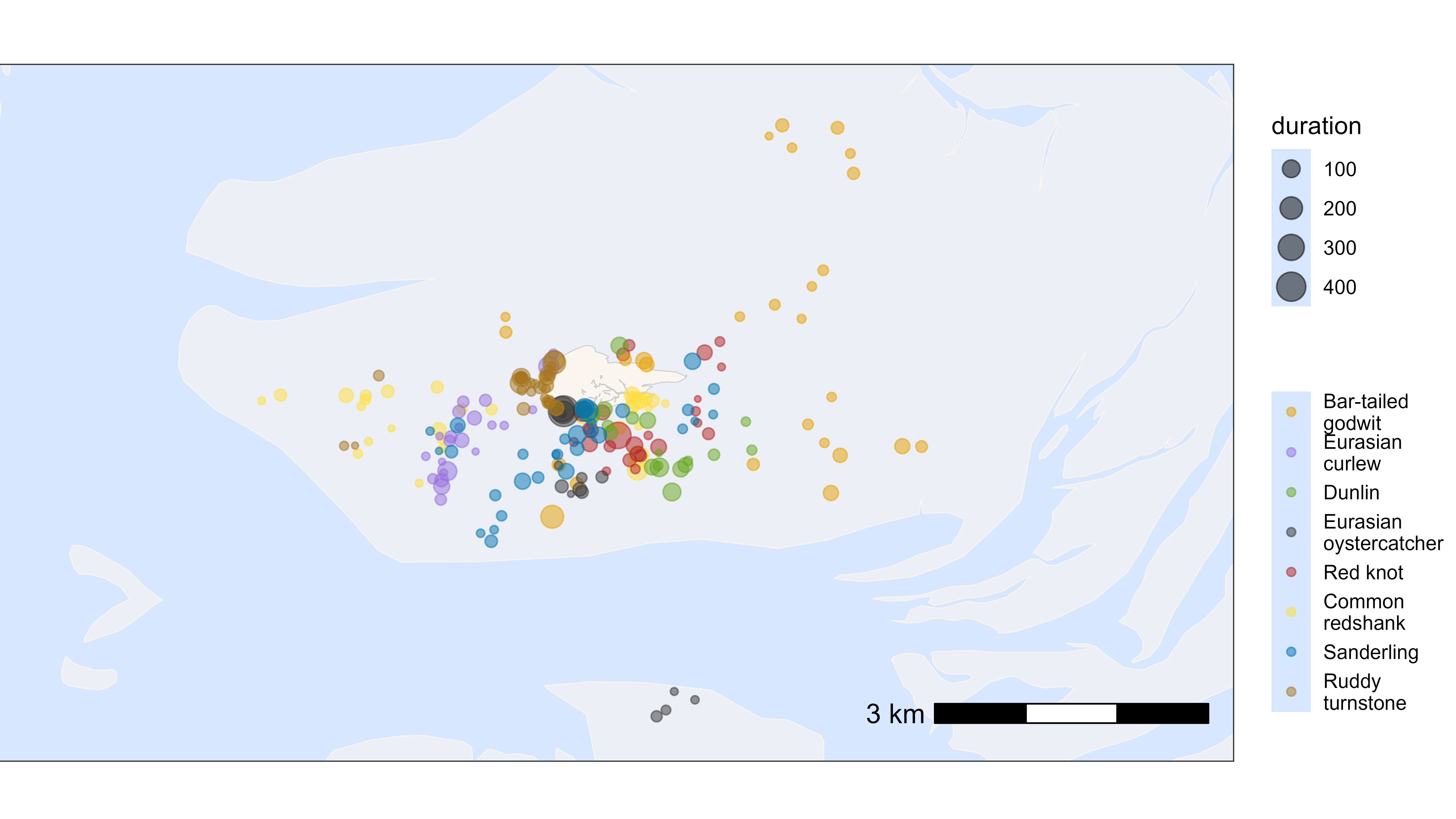
Plot by species
Similarly, we can plot the residence patches by species. For this, we need to merge the species information back to the summary table for residence patches. The residence patches are coloured by species and scaled by duration (in minutes).
# create basemap
bm <- atl_create_bm(data, buffer = 500)
# add species
du <- unique(data, by = "tag")
data_summary <- data_summary[du, on = "tag", `:=`(species = i.species)]
# plot residence patches itself by duration and species
bm +
geom_point(
data = data_summary,
aes(x_median, y_median, color = species, size = duration),
show.legend = TRUE, alpha = 0.5
) +
scale_color_manual(
values = atl_spec_cols(),
labels = atl_spec_labs("multiline"),
name = ""
)